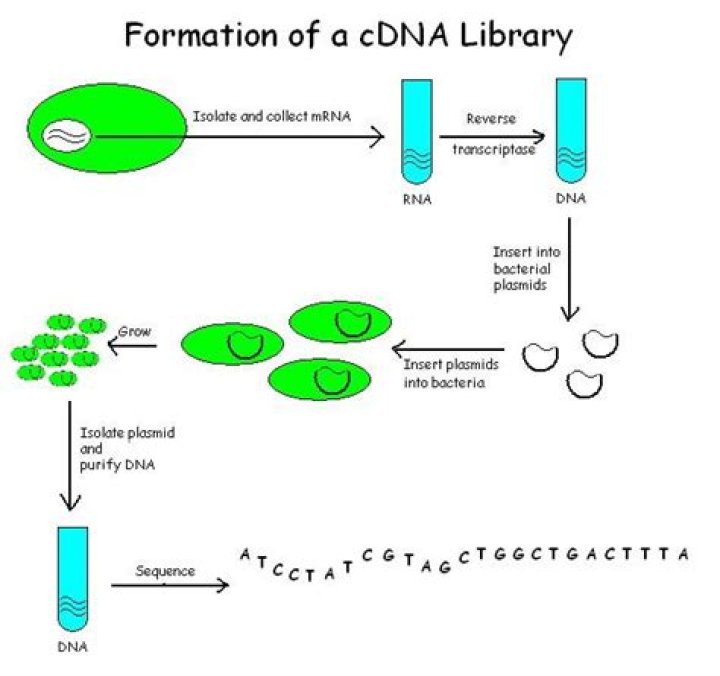
How is a cDNA library made

Robert Spencer
Published Apr 02, 2026
A cDNA library represents a collection of only the genes that are encoded into proteins by an organism. Complementary DNA, or cDNA, is created through reverse transcription of messenger RNA, and a library of cDNAs is generated using DNA cloning technology.
How is cDNA library constructed?
Creation of a cDNA library starts with mRNA instead of DNA. Messenger RNA carries encoded information from DNA to ribosomes for translation into protein. To create a cDNA library, these mRNA molecules are treated with the enzyme reverse transcriptase, which is used to make a DNA copy of an mRNA (i.e., cDNA).
How are cDNA libraries made quizlet?
How is a cDNA library created? isolating mRNA from a cell, then using reverse transcriptase to make cDNA. DNA polymerase is then used to make the cDNA double-stranded.
How do you make a cDNA library from RNA?
- Prepare sample. RNA serves as the template in cDNA synthesis. …
- Remove genomic DNA. Trace amounts of genomic DNA (gDNA) may be co-purified with RNA. …
- Select reverse transcriptase. …
- Prepare reaction mix. …
- Perform cDNA synthesis. …
- Prepare sample. …
- Remove genomic DNA. …
- Select reverse transcriptase.
How is cDNA created?
In cellular life, cDNA is generated by viruses and retrotransposons for integration of RNA into target genomic DNA. In molecular biology, RNA is purified from source material after genomic DNA, proteins and other cellular components are removed. cDNA is then synthesized through in vitro reverse transcription.
What is in a cDNA library?
A cDNA library is a combination of cloned cDNA (complementary DNA) fragments inserted into a collection of host cells, which constitute some portion of the transcriptome of the organism and are stored as a “library”. … Similarly, tissue-specific cDNA libraries can be produced.
How do you build a DNA library?
DNA libraries are constructed by partially cutting the genome of interest with a restriction enzyme to generate large fragments, inserting each of the fragments into a vector, and then putting each vector into a bacterial cell. Each bacterium in a library has a different part of the genome.
How do I clone a cDNA library?
The most frequently used technique for cloning cDNAs involves the addition of complementary homopolymeric tracts to double stranded cDNA and to the plasmid vector. To the cDNA, strings of cytosine residues are added using the enzyme terminal transferase to form oligo-dC tails on the 3′ ends.What is the difference between genomic library and cDNA library?
The key difference between cDNA and Genomic library is that cDNA library contains the cloned complementary DNA of total mRNA of an organism while the genomic DNA library contains the cloned fragments of the entire genome of an organism. The genomic DNA library is larger than the cDNA library.
Can a cDNA be created from a protein?Some vectors are designed to express only the cDNAs, while others have been modified to express not only the cDNA but also to express it in a context so the cDNA is made into a protein or a fusion protein.
Article first time published onWhich is the source of cDNA library quizlet?
A cDNA library is made from mRNA sequences. Cellular mRNAs are isolated and then reverse transcriptase is used to copy the mRNA sequences to cDNA, which are cloned into plasmid or phage vectors.
What is cDNA library quizlet?
cDNA Library. Only has genes from an organism which are expressed at certain times, without introns. The cDNA is then stored in either bacteria or a phage, for easy access. You just studied 3 terms! 1/3.
What is the purpose of a cDNA library quizlet?
What is the purpose of a cDNA library? To produce a library of expressed genes.
What is a cDNA library and what is its function in genetics?
cDNA library is a collection of the complementary DNA that is extracted from a cloned organism.
What does a DNA library consists of?
A DNA library is a collection of DNA fragments that have been cloned into vectors so that researchers can identify and isolate the DNA fragments that interest them for further study. There are basically two kinds of libraries: genomic DNA and cDNA libraries.
What are three key differences between a genomic and a cDNA library?
Genomic LibrarycDNA librariesIt is largerIt is smallerIt represents the entire genome of an organism having both coding and non coding regions.It represents only the expressed part of the genome and contain only coding sequences called ESTs
How is genomic DNA library constructed?
In order to construct a genomic library, the organism’s DNA is extracted from cells and then digested with a restriction enzyme to cut the DNA into fragments of a specific size. The fragments are then inserted into the vector using DNA ligase. … Genomic libraries are commonly used for sequencing applications.
Why do scientists prefer cDNA library over genomic library?
There are several advantages to using cDNA as opposed to genomic DNA for doing this: No introns: Eukaryote genes commonly contain introns (non-coding sequences). … More template: There are multiple copies of mRNA for every copy in the genome, so means you will get more copies per cell of the sequence of interest.
What are the limitations of cDNA library?
The disadvantage of a cDNA library is that it contains only sequences that are present in mature mRNA. Introns and any other sequences that are altered after transcription are not present; sequences, such as promoters and enhancers, that are not transcribed into RNA also are not present in a cDNA library.
What are the two main differences between the screening processes of genomic and cDNA libraries?
- The cDNA clone will only contain the sequences found in the mRNA, not the entire gene while the genomic clone could have the sequences of the entire gene.
- A cDNA library will not contain a clone of every gene of the organism.
Which of the following is most necessary to make cDNA library?
(g) The following enzymes are needed to make a cDNA library: DNA polymerase, DNA ligase, RNA polymerase, restriction enzymes.
Does cDNA have promoter?
cDNA does not contain any promoter sequences. In addition, because it was made from mRNA, all the introns have been removed. … The cloning vector would need to have a promoter that bacterial RNA polymerase recognizes and uses for transcription.
Can cDNA be patented?
New Supreme Court Decision Rules That cDNA Is Patentable What It Means for Research and Genetic Testing. In a unanimous decision last month, the Supreme Court ruled that naturally occurring genes are not patentable. But, said the Court, cDNA, a man-made copy of the genetic messenger in cells, is patentable.
How does the construction of a genomic DNA library differ from that of a cDNA library quizlet?
A) A genomic library contains both noncoding sequences and coding sequences, whereas a cDNA library contains only coding sequences.
How do bacteria protect their own DNA from restriction enzymes quizlet?
Restriction enzymes cut foreign DNA, such as viral DNA, into fragments. Bacteria protect their own DNA by modifying bases, usually by methylation, at the recognition sites. … When an electric field is applied, the negatively charged DNA molecules migrate toward the positive electrode.
What is the function of the Crispr CAS system in nature?
CRISPR-Cas9 was adapted from a naturally occurring genome editing system in bacteria. The bacteria capture snippets of DNA from invading viruses and use them to create DNA segments known as CRISPR arrays. The CRISPR arrays allow the bacteria to “remember” the viruses (or closely related ones).
What is a genomic library quizlet?
A genomic library is a collection of the total genomic DNA from a single organism. The DNA is stored in a population of identical vectors, each containing a different insert of DNA.
What can terminal transferase be used for quizlet?
4- an enzyme called terminal transferase can be used to add dGTP nucleotides to the 3′ end tail.
What is a cDNA molecule?
Complementary DNA (cDNA) is a DNA copy of a messenger RNA (mRNA) molecule produced by reverse transcriptase, a DNA polymerase that can use either DNA or RNA as a template. From: Encyclopedia of Genetics, 2001.
Where did scientists discover the reverse transcriptase enzyme?
Reverse transcriptases were discovered by Howard Temin at the University of Wisconsin–Madison in Rous sarcoma virions and independently isolated by David Baltimore in 1970 at MIT from two RNA tumour viruses: murine leukemia virus and again Rous sarcoma virus.
Which type of DNA library represents the genes expressed by a given cell at a certain time?
Which type of DNA library represents the genes expressed by a given cell at a certain time? cDNA libraries represent expressed genes from specific cell types under specific conditions.